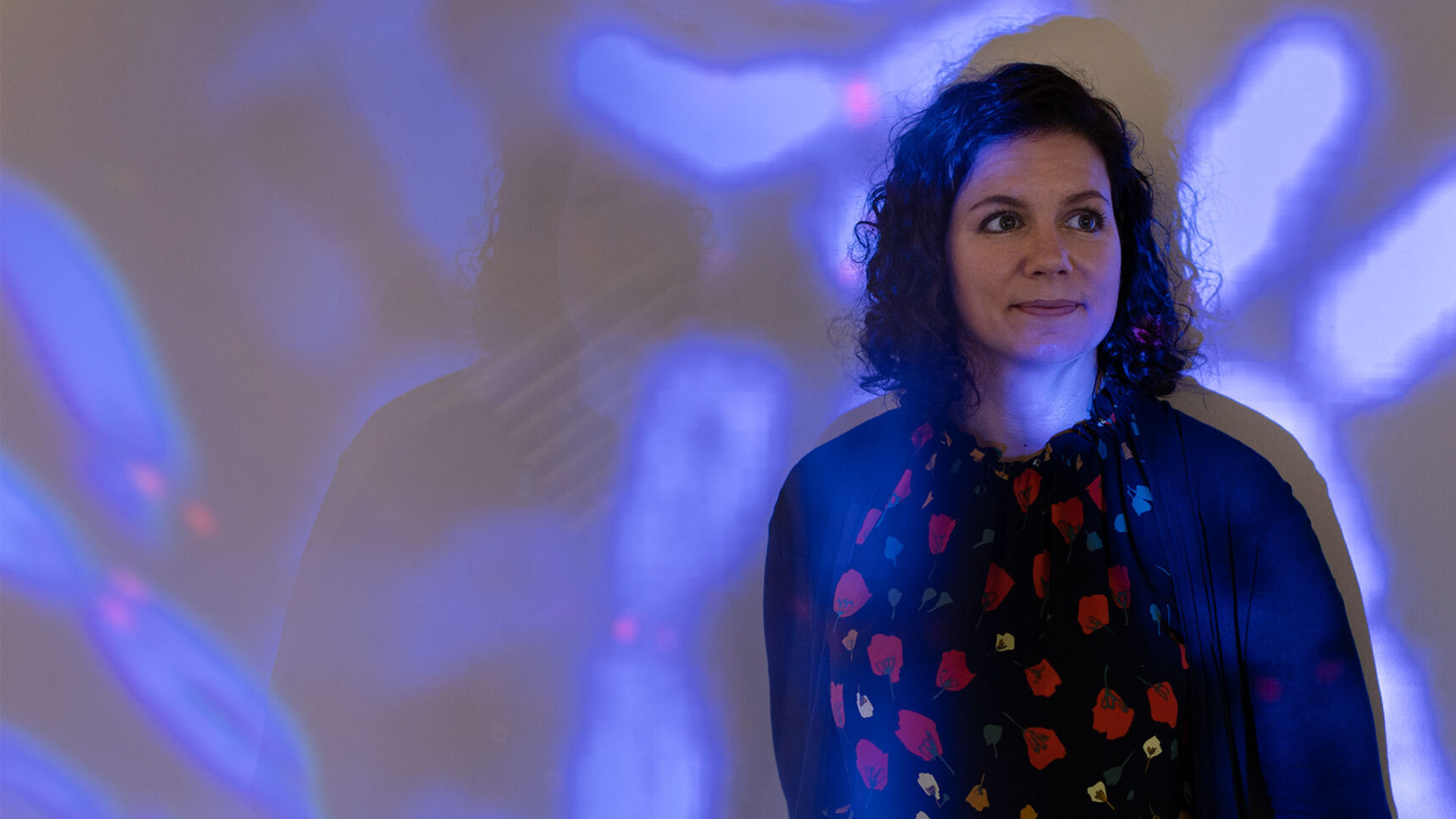
Karen Miga Fills In the Missing Pieces of Our Genome
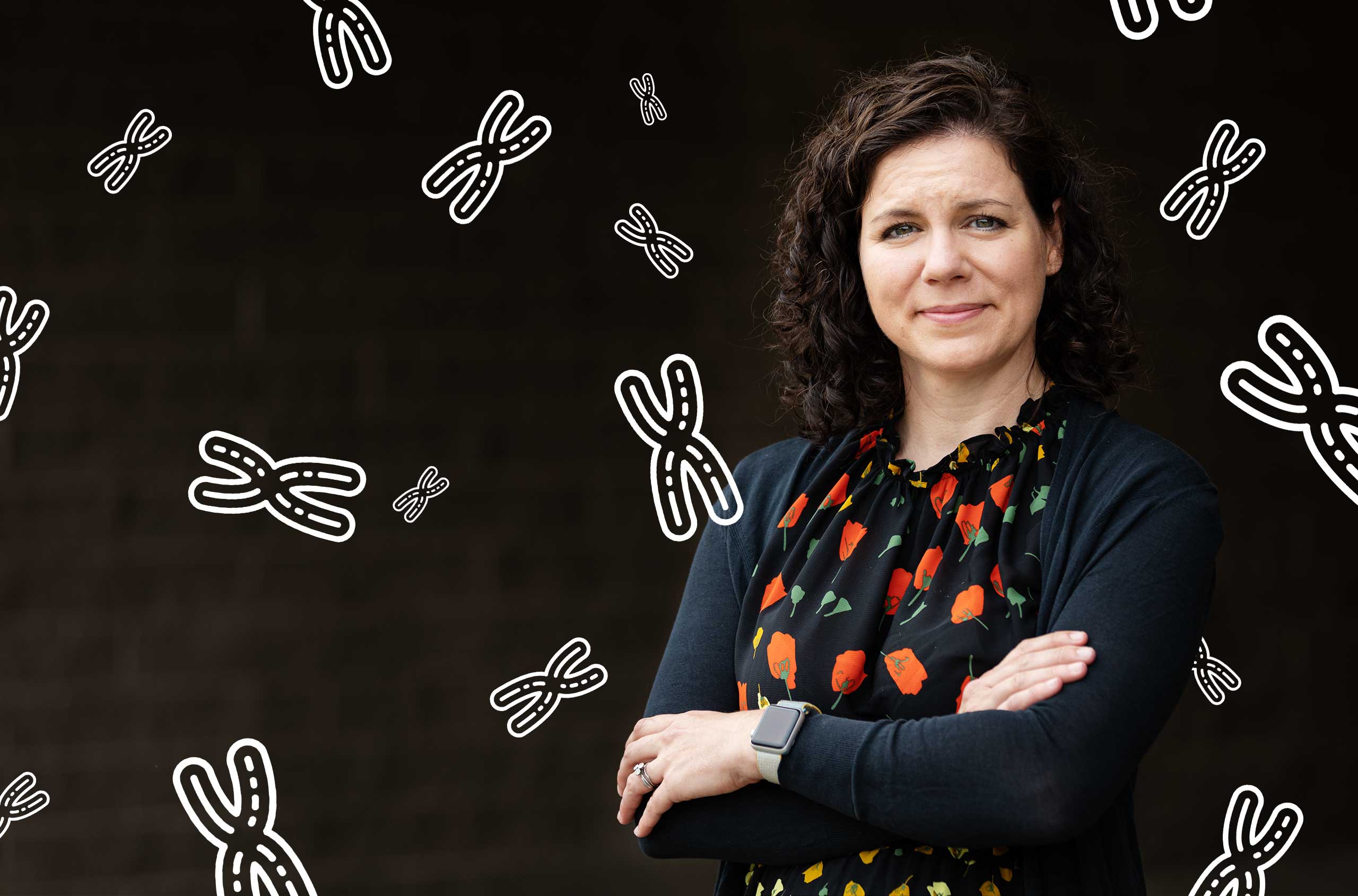

The geneticist Karen Miga, a leader of the Telomere-to-Telomere sequencing consortium, is intrigued by vital regions of our DNA that defied earlier efforts.
Photo-illustration: Quanta Magazine;
photo: Constanza Hevia for Quanta Magazine
Introduction
In 1990, an international team of scientists began an ambitious attempt to sequence the human genome. By 2001 the Human Genome Project (HGP) had prepared a rough draft, and in April 2003, the draft sequence was declared finished. But Karen Miga, a geneticist now at the University of California, Santa Cruz and the associate director of the UCSC Genomics Institute, knew that while the work might have wrapped up, the sequencing was far from complete.
The HGP was able to sequence the 90% of human DNA that geneticists call euchromatin, which is loosely folded and contains nearly all of the genes that are actively making proteins. But Miga specialized in heterochromatin, the tightly packed sections of DNA with highly repetitive sequences near the ends (telomeres) and centers (centromeres) of chromosomes. At the time, scientists couldn’t sequence heterochromatin, so despite the celebratory hubbub and champagne toasts, almost 10% of the genome went unsequenced.
It stayed that way for almost 20 years. The problem nagged at Miga, in part because she didn’t believe that the regions were as unimportant as some geneticists thought. (Without a sequence, how could you tell?) Over the years, Miga continued to push the genomics field to complete the project they had started so many years before. As DNA sequencing technologies enabled researchers to read longer and longer stretches of the genome in one go, Miga could see that scientists were inching closer to the possibility of cracking the problem open.
Together with Adam Phillippy, a computational biologist at the National Human Genome Research Institute, Miga launched the Telomere-to-Telomere (T2T) consortium in 2018 to finally sequence every last nucleotide of human DNA. Then, just as the team was finding its footing, the pandemic struck.
But COVID-19 didn’t stop their progress. In June, Miga, Phillippy and their colleagues published the first complete genome sequence on the preprint server biorxiv.org. Three decades after it began, the human genome was finally complete.
Quanta sat down with Miga in a video chat to discuss her years of work and what the consortium’s accomplishment might mean for science. The interview has been condensed and edited for clarity.
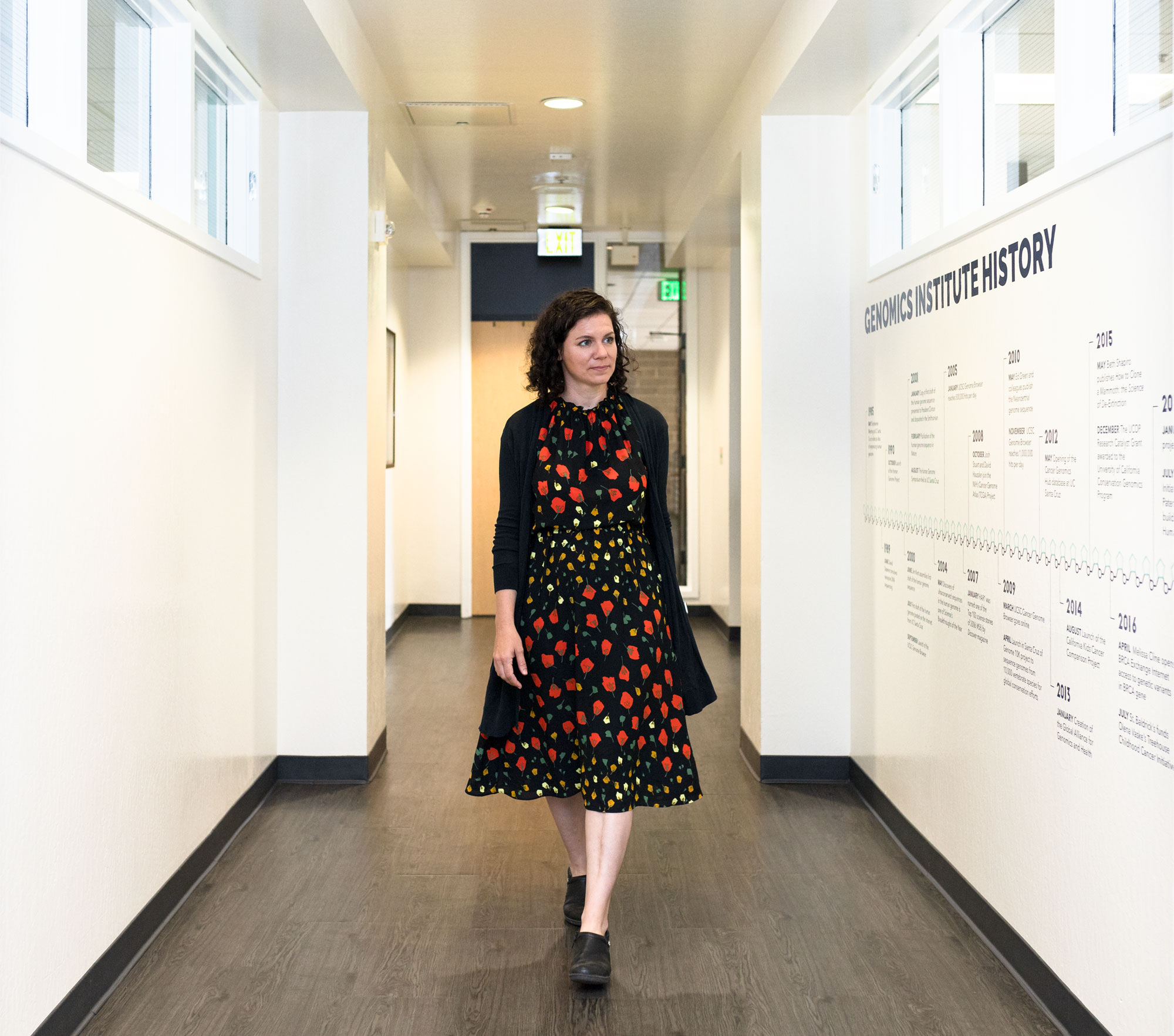

“It’s only now that we can look at ‘long read’ sequences that we can start to make accurate maps,” Miga explained.
Constanza Hevia for Quanta Magazine
How did you first become interested in genomics?
I was first introduced to human genomics and repeat biology at Case Western Reserve, when I was a master’s student in Evan Eichler’s lab. That was when the “completion” of the Human Genome Project was being announced, and because he was a leader in that field and very invested in understanding complex regions, I was well aware from the start that the genome was incomplete. Then I completed my doctoral work with Huntington Willard, a leader in centromere genomics and chromosome biology, at Duke University. It was under Hunt’s mentorship that I found my love for satellite DNA (sequence repeats that are found in tandem, or a head-to-tail arrangement, often for millions of bases), and I haven’t turned back.
What is satellite DNA, and what does it have to do with why the human genome remained unfinished?
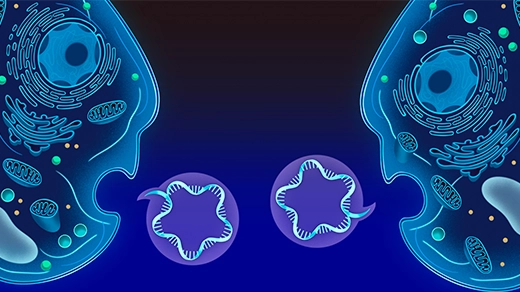
Short tandem repeats are common throughout the human genome and have been well-studied. The satellite DNAs that I was most passionate about were quite different in both scale and function in genome biology. In terms of scale, rather than being interspersed in gene-rich regions, these satellites DNAs constitute extremely large, gene-poor regions (often tens of millions of bases) on every human chromosome. We know that these regions enriched in satellite DNAs are a common feature of plant and animal genomes. Further, we know that they are super important for cell viability. They mark sites of centromere formation — parts of our genome that ensure that our chromosomes segregate correctly during cell division. These incredibly strange genomic landscapes were missing from our earlier assemblies of the human genome. And of course, they were also missing from the studies that used those assemblies to make new genetic discoveries and associations with human diseases.
What parts of the human genome remained unsequenced at that point?
The biggest gaps that remained in the human genome were all repetitive DNA. In the case of satellite DNAs, the tandem repeats are organized in a linear array of near exact copies. You may have a single nucleotide change that could distinguish one copy from another, and those differences may be spaced apart by tens of thousands bases. If you could only sequence 150 bases at a time, you never would have enough information in that stretch of letters to tell where in the genome that segment is from. In the past, when we used small fragments like that, we couldn’t use them to fully resolve these regions that are extremely rich in repeats. It’s only now that we can look at “long read” sequences that we can start to make accurate maps.
So to finish the sequencing, was it only a matter of technological improvement?
Our team absolutely benefitted from the availability of long-read sequencing. However, another major consideration here is the technological advancement in developing the right algorithms that use these long reads to make the best linear predictions. Additionally, our T2T community lead the development of new analyses to evaluate the quality of these predictions to ensure that they are accurate.
I understand the desire to finish the full human genome sequence just to have it done. But what were we missing by leaving these genomic segments unsequenced?
There’s merit in having a more complete assessment of the total number of genes in a human genome. Our preprint identifies hundreds of protein-coding genes and over a thousand gene predictions. That’s an important addition to the human genome and provides clear merit to the work.
Apart from genes, it is important to remember that we are releasing sequences for entire short arms of our chromosomes. For example, having three copies of chromosome 21 (trisomy 21) is of clear clinical interest, because that leads to Down syndrome. We released the first map of the short arm of chromosome 21, which adds information about the genomic organization of that chromosome that could broaden functional and clinical studies. Additionally, we now have a clear view of functionally important regions, such as centromeres, and that could help us launch new studies to better understand their genome organization and structure.
When the draft genome was published in 2001, how much was actually finished?
The very first draft sequence was an incredible resource, but it intentionally ignored the more complex, densely repetitive regions of our genome. Later efforts to get to a more finished state had about 8-10% still missing, and we kept hitting a technological wall to reach completion. In part this was due challenges in sequencing the repeats, but it is important to remember that even if the sequencing was perfect, we would still be faced with the obstacle of correctly putting those pieces together.
I feel like I am one of a small group of scientists who have been standing on our soapboxes for many years, saying, “Hey, our current maps are incomplete, and completing our maps will be important to our understanding of genome biology.” I suspect that many will be surprised to learn this, due to the very public celebration of the “finished” human genome in 2003. We were celebrating the completion of the parts of our genome that were technologically feasible at the time. It wasn’t really clear to the public how much of our genome was left unresolved and unexplored.

The centromeres, where replicating chromosomes attach to each other and to spindle fibers during cell division, are rich in extremely repetitive “satellite DNA” sequences.
Constanza Hevia for Quanta Magazine
You helped head up the T2T consortium to map these long, repetitive parts of the human genome. How did the T2T consortium get started?
Adam Phillippy at the National Human Genome Research Institute (NHGRI) and I were both interested in completing a human genome, and we started working together in 2018. Initially, we started with the goal of releasing one complete “telomere-to-telomere” human chromosome. The public announcement of the T2T consortium happened in 2019 at a meeting for Advances in Genome Biology and Technology, when Adam presented our work to complete the X chromosome and announced the launch of our initiative to complete a human genome.
Following our effort to complete a single chromosome, we continued to study how to improve chromosome assemblies using combinations of both highly accurate data and ultra-long reads. We started to reach more automated methods to complete the satellite arrays. Through it all, the consortium was growing and transforming into a broader grassroots effort with a huge cast of talented scientists.
In the summer of 2020, we formed a focused workshop to try and finish all of the human genome. This was key to organizing our virtual team into distinct working groups, each with dedicated expertise in assembly, curation, variant calling and repeat biology. This virtual community worked together — notably, throughout a global pandemic — to offer an initial release of a complete human genome, along with a large number of in-depth biological analyses from these newly assembled regions.
What were some of the challenges you faced along the way?
We had several challenges. A notable one was a long array of satellite DNA next to the centromere on chromosome 9, which had a large duplication that took real effort to resolve. Adam Phillippy and his team at NHGRI deserve special credit for their focused work to resolve the sequences of the ribosomal DNA arrays on each of the five acrocentric chromosomes, where the centromere is much closer to one end than to the center.
How did you feel when you finally had the whole genome assembly?
It’s a dream. This is really like waking up to a dream come true. I had always fantasized about having these types of maps when I was a graduate student, and I always thought that they would arrive someday. I’m just so grateful to be part of the process of issuing and sharing this information.
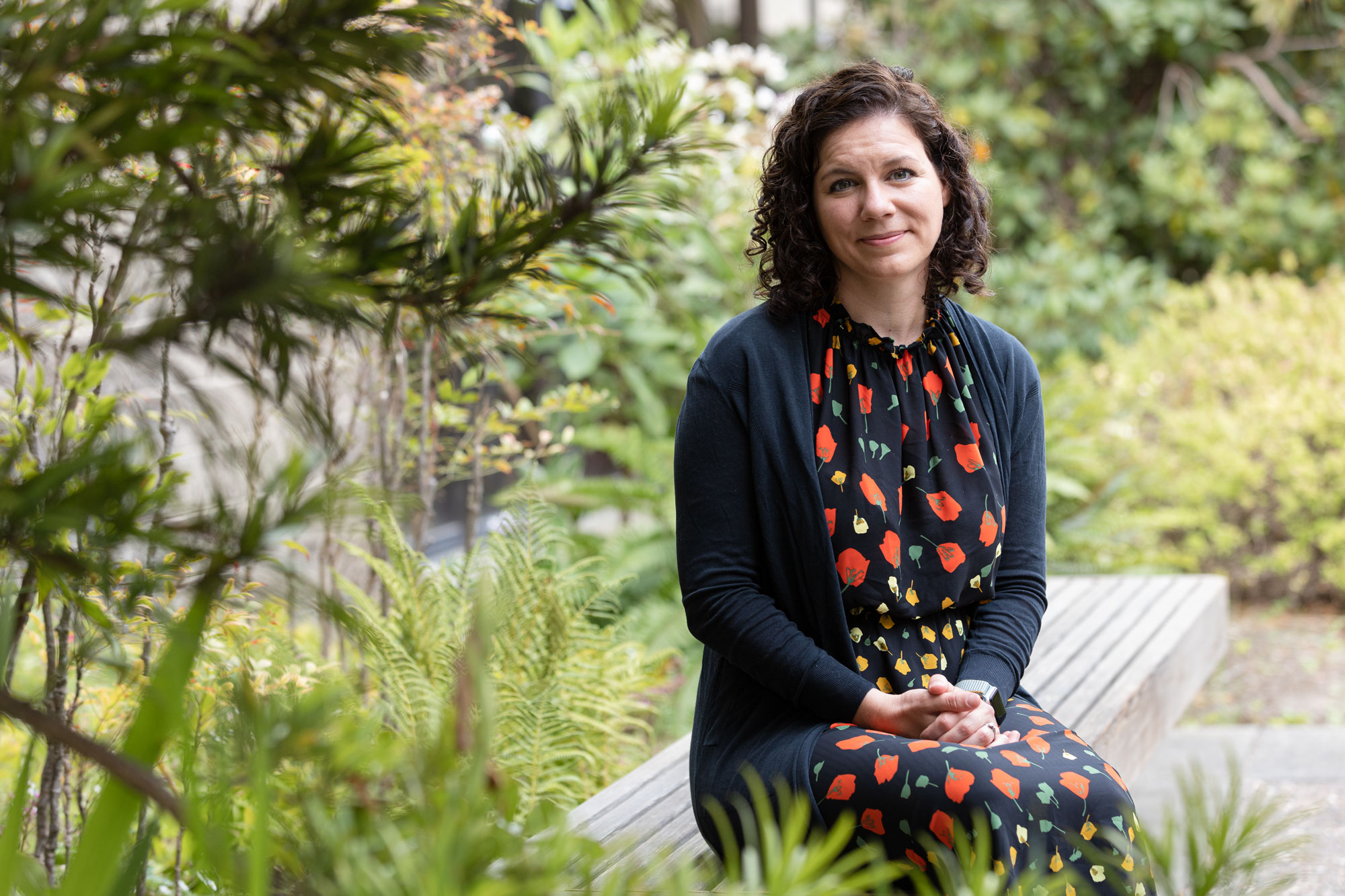

“I’m just so grateful to be part of the process of issuing and sharing this information,” Miga said.
Constanza Hevia for Quanta Magazine
What does it mean to have finally completed sequencing the human genome?
When the first complete human genome is published, it will be a landmark moment. It will be a huge technological achievement in that it will be the first time we will have demonstrated this new standard of genome completion and quality. It will also be a huge win for the broader basic and translational research communities, as we will have hundreds of millions of new, unexplored bases in a human genome to expand studies of disease association and cellular function. The next challenge is to make this completely routine and expand our study across hundreds if not thousands of people.
To reach this next step, the T2T Consortium formally partnered with the Human Pangenome Reference Consortium in 2020, with a common goal to reach complete, finished assemblies of hundreds of diverse human genomes. I would hope that the sequencing technology and assembly process could be more streamlined in the future to ensure that reaching a T2T genome becomes standard operating procedure.
What are some of the basic science questions the complete human genome sequence will enable scientists to explore?
Sequences in these regions have their own specialized models and rates of evolution, and they offer new genetic information to help us understand genome structure and function. We have essentially opened the door to studying many gene families, entire acrocentric chromosomes and new noncoding regions of the human genome in greater detail. My lab is very interested in studying how satellite DNAs vary between diverse individuals and how this new source of sequence variation contributes to our understanding of genome biology and human disease.
Yet by fully sequencing the human genome, you’ve also opened the door to understanding other important mechanisms of genetics.
Centromeres and other regions that are repeat rich regions are present in our maps for the first time. This offers a largely unexplored genomic landscape for new discoveries and it can expand genetic studies of disease association. Early work suggests that these parts of the genome are organized, replicated and regulated differently than the more familiar gene-rich regions we have intensively studied since the initial release of the human reference genome.
So it’s not like the T2T preprint genome is just topping off the previous human genome reference with a last few details. It’s more like there’s a whole chunk of our genome that operates in a different way just becoming available to us, and we have only begun to scratch the surface of that. The next 10 years should be very exciting, and I look forward to the future discoveries in these newly revealed regions.